309.2. Filter Transformations#
309.2. Filter transformations¶
For the Rubin Science Platform at data.lsst.cloud.
Data Release: Data Preview 1
Container Size: medium
LSST Science Pipelines version: v29.2.0
Last verified to run: 2026-03-24
Repository: github.com/lsst/tutorial-notebooks
DOI: 10.11578/rubin/dc.20250909.20
Learning objective: An overview of transformation relations to and from the LSSTComCam photometric system presented in Data Preview 1 and other commonly-used photometric systems and how to apply these relations.
LSST data products: Object
Packages: lsst.rsp, lsst.afw.display
Credit: Originally developed by the Rubin Community Science team. Please consider acknowledging them if this notebook is used for the preparation of journal articles, software releases, or other notebooks.
Get Support: Everyone is encouraged to ask questions or raise issues in the Support Category of the Rubin Community Forum. Rubin staff will respond to all questions posted there.
1. Introduction¶
Being able to transform measurements between different photometric systems is essential for comparing results with not only contemporaneous and legacy data sets but also with theoretical models (e.g., stellar isochrones) derived using legacy photometric systems.
To this end, the Vera C. Rubin Project has generated transformation relations between the Data Preview 1 (DP1) $ugrizy$ system and other systems, like PanSTARRS1, Dark Energy Survey (DES), EUCLID, Gaia DR3, Johnson-Cousins, and others. These transformation relations are provided in Rubin Technical Note #99 (RTN-099). (For those interested, Jupyter notebooks used for calculating the transformation relations provided in RTN-099 can be found in this github repository.)
Two methods for transforming between photometric systems are explored in RTN-099: polynomial relations and lookup tables. The guiding principle for which is to use is driven by the balance between simplicity and accuracy: in general, simple first- or second-order polynomials are sufficient, but, for higher accuracy, one may instead choose the slightly more difficult-to-employ lookup-table method.
This tutorial demonstrates how to apply both methods of photometric transformation. Examples are provided for converting from Rubin DP1 $i$-band photometry to PanSTARRS1 DR2 and DES DR2 $i$ bands, and from Rubin DP1 $g$-band photometry to Gaia DR3 $BP$-band photometry.
Related tutorials: The 100-level tutorials demonstrate how to use the butler, the TAP service, and the Firefly image display. The 200-level tutorials introduce the types of image and catalog data.
1.1. Import packages¶
Import general python packages math, numpy, pandas, scipy, astropy, and matplotlib.
Import the pyvo module for accessing remote data.
From the lsst package, import the module for accessing the Table Access Protocol (TAP) service.
import math
import numpy as np
import pandas as pd
from scipy import interpolate
from astropy.coordinates import SkyCoord
import astropy.units as u
import matplotlib.pyplot as plt
import pyvo
from lsst.rsp import get_tap_service
1.2. Define parameters and functions¶
Create an instance of the TAP service, and assert that it exists.
service = get_tap_service("tap")
assert service is not None
Define the approximate central coordinates of the ECDFS field and a radius of 0.25 degrees about this center.
Note: The radius is restricted to 0.25 degrees here, since the public PanSTARRS1 DR2 TAP service -- which is used later in this notebook -- is currently restricted to cone searches of no larger than 0.25 degrees.
ra_cen = 53.2
dec_cen = -28.1
radius = 0.25
Define parameters to use colorblind-friendly colors with matplotlib.
plt.style.use('seaborn-v0_8-colorblind')
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
Define a function for cross-matching between sky catalogs.
def cross_match_catalogs(df1, df2, ra_name_1, dec_name_1, ra_name_2, dec_name_2):
"""Match two catalogs based on RA, Dec coordinates.
Use a fixed, 3-arcsecond matching radius, keeping only the closest
match in cases where there is more than one.
Parameters
----------
df1 : `DataFrame`
First catalog to be matched.
df2 : `DataFrame`
Second catalog to be matched.
ra_name_1 : `string`
Name of the column in df1 containing the right ascension.
dec_name_1 : `string`
Name of the column in df1 containing the declination.
ra_name_2 : `string`
Name of the column in df2 containing the right ascension.
dec_name_2 : `string`
Name of the column in df1 containing the declination.
Returns
-------
matches : `DataFrame`
Table of matched objects.
unmatched : `DataFrame`
Table of unmatched objects from df1.
"""
coords1 = SkyCoord(ra=df1[ra_name_1].values*u.degree,
dec=df1[dec_name_1].values*u.degree)
coords2 = SkyCoord(ra=df2[ra_name_2].values*u.degree,
dec=df2[dec_name_2].values*u.degree)
max_sep = 3 * u.arcsec
idx, d2d, d3d = coords1.match_to_catalog_sky(coords2)
mask = d2d < max_sep
valid_idx_mask = idx[mask] < len(df2)
combined_mask = mask.copy()
combined_mask[mask] = valid_idx_mask
matches = df1[combined_mask].copy()
matches['match_idx'] = idx[combined_mask]
matches['separation_arcsec'] = d2d[combined_mask].arcsec
for col in df2.columns:
matches[f'match_{col}'] = df2.iloc[idx[combined_mask]][col].values
matches = matches.loc[matches.groupby(matches.index)['separation_arcsec'].idxmin()]
unmatched = df1[~combined_mask]
return matches, unmatched
2. Grab LSSTComCam ECDFS point source data from DP1¶
Define a query to grab stars in the ECDFS field from the DP1 Object table. To avoid potentially saturated stars and very faint stars, restrict the sample returned to those point sources with $r$-band magnitudes between 16.0 and 24.0.
query = """SELECT coord_ra, coord_dec, objectId,
u_psfMag, g_psfMag, r_psfMag, i_psfMag, z_psfMag,
u_psfMagErr, g_psfMagErr, r_psfMagErr, i_psfMagErr, z_psfMagErr,
refExtendedness
FROM dp1.Object
WHERE CONTAINS(POINT('ICRS', coord_ra, coord_dec), CIRCLE('ICRS', %f, %f, %f))=1
AND refExtendedness=0
AND r_psfMag BETWEEN 16.0 AND 24.0""" % (ra_cen, dec_cen, radius)
Submit asychronous query.
job = service.submit_job(query)
job.run()
job.wait(phases=['COMPLETED', 'ERROR'])
print('Job phase is', job.phase)
if job.phase == 'ERROR':
job.raise_if_error()
Job phase is COMPLETED
Put results into a pandas dataframe.
df_dp1 = job.fetch_result().to_table().to_pandas()
print(f"Total number of visits: {len(df_dp1)}")
Total number of visits: 979
Display table.
df_dp1
| coord_ra | coord_dec | objectId | u_psfMag | g_psfMag | r_psfMag | i_psfMag | z_psfMag | u_psfMagErr | g_psfMagErr | r_psfMagErr | i_psfMagErr | z_psfMagErr | refExtendedness | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 53.426710 | -28.048048 | 611254316728079110 | 20.344200 | 18.753799 | 18.169399 | 17.973700 | 17.897800 | 0.003464 | 0.000244 | 0.000190 | 0.000247 | 0.000319 | 0.0 |
| 1 | 53.448204 | -28.037606 | 611254316728079756 | 25.531500 | 23.614901 | 22.276501 | 21.103300 | 20.583799 | 0.339714 | 0.008660 | 0.003279 | 0.002096 | 0.002466 | 0.0 |
| 2 | 53.451077 | -28.040253 | 611254316728079755 | 20.675301 | 18.126600 | 16.837200 | 15.779200 | 15.337100 | 0.004685 | 0.000182 | 0.000100 | 0.000092 | 0.000093 | 0.0 |
| 3 | 53.463196 | -28.035149 | 611254316728079531 | 17.867901 | 16.735001 | 16.342600 | 16.230000 | 16.196600 | 0.000819 | 0.000100 | 0.000082 | 0.000112 | 0.000124 | 0.0 |
| 4 | 53.414971 | -28.053233 | 611254316728078899 | 26.046600 | 23.978100 | 22.803400 | 22.283100 | 22.035400 | 0.492280 | 0.011412 | 0.005248 | 0.005710 | 0.008755 | 0.0 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 974 | 53.388366 | -28.254669 | 611253629533308555 | 18.524000 | 17.586901 | 17.209499 | 17.072901 | 17.031099 | 0.001083 | 0.000132 | 0.000117 | 0.000146 | 0.000192 | 0.0 |
| 975 | 53.382798 | -28.251805 | 611253629533309059 | 25.473801 | 23.990400 | 22.610901 | 21.828899 | 21.486099 | 0.311844 | 0.011550 | 0.004326 | 0.003659 | 0.005661 | 0.0 |
| 976 | 53.312514 | -28.250762 | 611253629533308882 | 20.676399 | 19.047300 | 18.407200 | 18.164400 | 18.058201 | 0.004232 | 0.000268 | 0.000200 | 0.000235 | 0.000321 | 0.0 |
| 977 | 53.304486 | -28.244330 | 611253629533308881 | 23.137400 | 20.602600 | 19.289600 | 17.962799 | 17.385300 | 0.034628 | 0.000695 | 0.000330 | 0.000212 | 0.000207 | 0.0 |
| 978 | 53.073823 | -28.065055 | 611254454167030204 | 21.985500 | 19.752001 | 18.740499 | 18.337200 | 18.149000 | 0.012172 | 0.000387 | 0.000237 | 0.000262 | 0.000335 | 0.0 |
979 rows × 14 columns
Calculate color indices using the PSF magnitudes, and plot a color-color diagram. Color-code points by the $r$-band PSF magnitude error.
df_dp1['gr_psfMag'] = df_dp1['g_psfMag'] - df_dp1['r_psfMag']
df_dp1['ri_psfMag'] = df_dp1['r_psfMag'] - df_dp1['i_psfMag']
df_dp1['gi_psfMag'] = df_dp1['g_psfMag'] - df_dp1['i_psfMag']
plt.figure(figsize=(10, 6))
sc = plt.scatter(
df_dp1['gr_psfMag'],
df_dp1['ri_psfMag'],
c=df_dp1['r_psfMagErr'],
cmap='viridis',
alpha=0.7,
s=8,
vmin=0.00,
vmax=0.02
)
plt.xlabel('g - r')
plt.ylabel('r - i')
plt.title('color-color diagram for 16<r<24 point sources in ECDFS')
plt.grid(True)
plt.xlim(-0.5, 2.0)
plt.ylim(-0.5, 2.5)
cbar = plt.colorbar(sc)
cbar.set_label('r_psfMagErr')
plt.show()
Figure 1: ($r$-$i$) vs. ($g$-$r$) color-color diagram for point sources in a 0.25-degree radius of the ECDFS field in DP1 data. Points are calculated from
Objecttable PSF magnitudes, and are color-coded by their $r$-band PSF magnitude error, ranging from 0 to 0.02 magnitudes. The figure shows the typical stellar locus as a narrow sequence with a sharp upturn at red colors.
3. First-order polynomial transformation: LSSTComCam $i$ --> PanSTARRS1 DR2 $i$¶
As a simple first case, look at the polynomial transformation from DP1 $i$-band to PanSTARRS1 DR2 $i$-band.
| Conversion | Transformation Equation | RMS | Applicable Color Range | QA Plot |
|---|---|---|---|---|
| $i_{ComCam} \to i_{ps1}$ | $i_{ps1} - i_{ComCam} = +0.014 (g-i)_{ComCam} - 0.016$ | 0.01 | $-2.2 < (g-i)_{ComCam} \leq 3.0$ | link |
Here are the quality assurance (QA) plots from RTN-099 associated with the first-order polynomial fit of the DP1 $i$-band to PanSTARRS1 DR2 $i$-band:
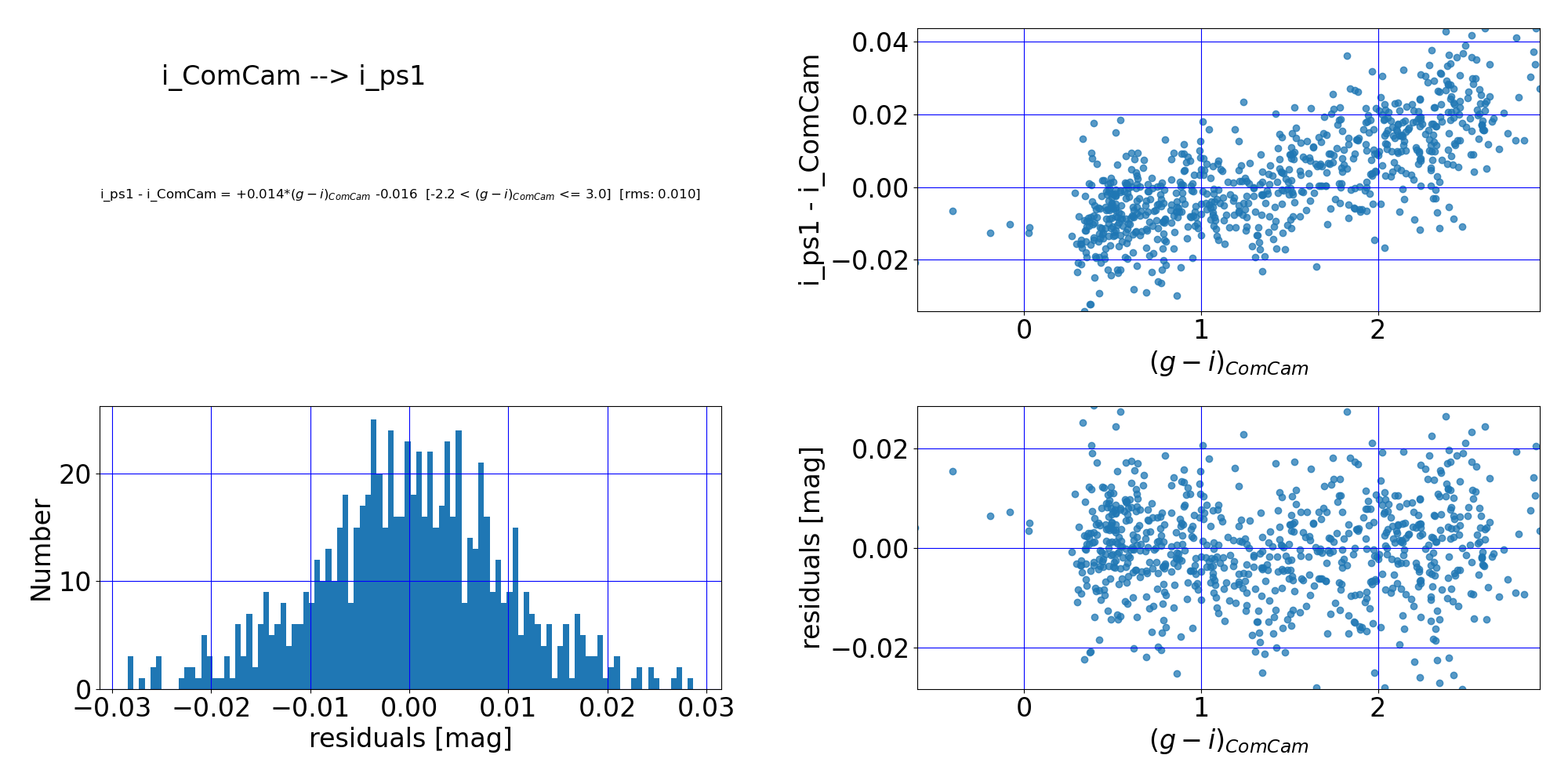
Figure 2: Comparison of LSSTComCam and PS1 DR2 $i$-band photometry. (Top left:) The fit polynomial equation, the RMS of the fit, and the applicable color range. (Top right:) The difference between the two magnitudes vs. color before any transformation. (Bottom left:) Histogram of the residuals of the fit. (Bottom right:) Residuals of the fit vs. ($g$-$i$) color. The residuals scatter about zero, suggesting that the fit has done a reasonable job.
3.2 Apply the transformation¶
Select only stars within the applicable color range (-2.2 < $g-i$ < 3.0) from the fit. Apply the polynomial transformation to these stars from the LSSTComCam data in the ECDFS field.
pick_ps1dr2_colors = (df_dp1['gi_psfMag'] < 3.0) & (df_dp1['gi_psfMag'] > -2.2)
df_dp1_ps1dr2_transform = df_dp1[pick_ps1dr2_colors].copy()
df_dp1_ps1dr2_transform['i_ps1_poly_offset'] = 0.014*df_dp1_ps1dr2_transform['gi_psfMag'] - 0.016
df_dp1_ps1dr2_transform['i_ps1_poly'] = df_dp1_ps1dr2_transform['i_psfMag'] +\
df_dp1_ps1dr2_transform['i_ps1_poly_offset']
# display(df_dp1_ps1dr2_transform)
3.3 Test the transformation¶
Test the transformation by comparing the PS1 $i$-band magnitudes estimated from the DP1 data in ECDFS against the PS1 $i$-band magnitudes from the PanSTARRS1 DR2 data themselves.
3.3.1 Grab ECDFS data from PanSTARRS1 DR2¶
Use pyvo to query the PanSTARRS1 DR2 TAP service, using suitable constraints for point sources. Since the PanSTARRS1 DR2 TAP service has a cone search maximum search radius of 0.25 deg, keep the search to that fixed radius.
ps1dr2_tap_url = 'https://mast.stsci.edu/vo-tap/api/v0.1/ps1dr2'
ps1dr2_tap = pyvo.dal.TAPService(ps1dr2_tap_url)
query = """
SELECT
o.objName,
o.raMean, o.decMean, o.raMeanErr, o.decMeanErr,
o.qualityFlag,
o.gMeanPSFMag, o.gMeanPSFMagErr, o.gMeanPSFMagNpt,
o.rMeanPSFMag, o.rMeanPSFMagErr, o.rMeanPSFMagNpt,
o.iMeanPSFMag, o.iMeanPSFMagErr, o.iMeanPSFMagNpt,
o.zMeanPSFMag, o.zMeanPSFMagErr, o.zMeanPSFMagNpt,
o.yMeanPSFMag, o.yMeanPSFMagErr, o.yMeanPSFMagNpt,
o.rMeanKronMag, o.rMeanKronMagErr,
o.nDetections, o.ng, o.nr, o.ni, o.nz,o.ny,
o.gFlags, o.gQfPerfect,
o.rFlags, o.rQfPerfect,
o.iFlags, o.iQfPerfect,
o.zFlags, o.zQfPerfect,
o.yFlags, o.yQfPerfect,
soa.primaryDetection, soa.bestDetection
FROM dbo.MeanObjectView o
LEFT JOIN StackObjectAttributes AS soa ON soa.objID = o.objID
WHERE CONTAINS(POINT('ICRS', RAMean, DecMean),CIRCLE('ICRS',%f,%f,0.25))=1
AND o.nDetections > 5
AND soa.primaryDetection>0
AND o.gQfPerfect>0.85 and o.rQfPerfect>0.85 and o.iQfPerfect>0.85 and o.zQfPerfect>0.85
AND (o.rmeanpsfmag - o.rmeankronmag < 0.05)
""" % (ra_cen, dec_cen)
result = ps1dr2_tap.run_sync(query)
df_ps1 = result.to_table().to_pandas()
# display(df_ps1)
/opt/lsst/software/stack/conda/envs/lsst-scipipe-10.1.0-exact/lib/python3.12/site-packages/pyvo/dal/query.py:341: DALOverflowWarning: Partial result set. Potential causes MAXREC, async storage space, etc.
warn("Partial result set. Potential causes MAXREC, async storage space, etc.",
3.3.2 Match the ECDFS data from LSSTComCam with the ECDFS data from PanSTARRS1 DR2¶
Use the cross_match_catalogs function defined above to match the two catalogs based on the stars' coordinates.
matches_dp1_ps1, unmatched_dp1_ps1 = cross_match_catalogs(df_dp1_ps1dr2_transform, df_ps1,
'coord_ra', 'coord_dec',
'ramean', 'decmean')
# display(matches_dp1_ps1)
3.3.3 Compare the LSSTComCam-estimated PS1 i-band magnitudes with the PanSTARRS1 DR2 i-band magnitudes¶
Calculate both the differences between the DP1 $i$-band magnitude and the PanSTARRS1 DR2 $i$-band magnitude ("before transformation") and the differences between DP1-estimated PS1 $i$-band magnitude and the PanSTARRS1 DR2 $i$-band magnitude ("after transformation"):
matches_dp1_ps1['dmag_i_cc_ps1'] = matches_dp1_ps1['i_psfMag'] -\
matches_dp1_ps1['match_imeanpsfmag']
matches_dp1_ps1['dmag_i_ps1_poly'] = matches_dp1_ps1['i_ps1_poly'] -\
matches_dp1_ps1['match_imeanpsfmag']
Plot the histogram of the $\Delta$mags (both before transformation and post transformation).
dmag_orig = matches_dp1_ps1.loc[:, 'dmag_i_cc_ps1']
dmag_new = matches_dp1_ps1.loc[:, 'dmag_i_ps1_poly']
dmag_min = -0.2
dmag_max = 0.2
if len(dmag_new) < 100:
plt.hist(dmag_orig, bins=10, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=10, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after transformation')
else:
plt.hist(dmag_orig, bins=100, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=100, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after transformation')
plt.xlabel('$\\Delta$mag [mag]')
plt.ylabel('Number')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"mag difference between LSSTComCam DP1 and PanSTARRS1 DR2 $i$-band")
plt.show()
Figure 3: Histogram comparing LSSTComCam and PS1 DR2 $i$-band photometry for matched stars; differences are calculated as $\Delta$mag = DP1 mag - PS1 mag. The orange histogram shows the difference before applying the transformation, demonstrating that there is a tail to negative $\Delta$mag values. The blue histogram is the result after applying the polynomial transformation. The transformed difference is centered on a magnitude residual of zero, as expected.
Create a scatter plot of the $\Delta$mags (both before transformation and post transformation) vs. color. Note that LSSTCamCom DP1 $i$-band and PanSTARRS1 DR2 $i$-band are fairly similar; so the transformation is relatively small.
color = matches_dp1_ps1.loc[:, 'gi_psfMag']
dmag_min = -0.2
dmag_max = 0.2
color_min = -0.5
color_max = 3.5
plt.scatter(color, dmag_orig, color=colors[2], alpha=0.50, label='before transformation')
plt.scatter(color, dmag_new, color=colors[5], alpha=0.50, label='after transformation', marker='*')
plt.axis([color_min, color_max, dmag_min, dmag_max])
plt.xlabel('$(g-i)_{ComCam}$')
plt.ylabel('$\\Delta$mag [mag]')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"$\Delta$mag between LSSTComCam DP1 and PanSTARRS1 DR2 $i$-band vs. color")
plt.show()
Figure 4: Magnitude differences between LSSTComCam and PS1 DR2 $i$-band photometry for matched stars plotted as a function of $(g-i)$ color. Orange points show the difference before applying the transformation, demonstrating that the tail to negative $\Delta$mag values mostly happens at redder $(g-i)$ colors. Blue stars are the results after applying the polynomial transformation. The transformed difference is centered on a magnitude residual of zero, as expected.
4. Piecewise 1st-order polynomial transformation: LSSTComCam $i$ --> DES DR2 $i$¶
Now look at the polynomial transformation from DP1 $i$-band to DES DR2 $i$-band. This is a little more complicated than the previous example, as there is a break in the relation; so this polynomial had to be fit piece-wise, with a break at $(g-i)_{ComCam} = 1.8$.
| Conversion | Transformation Equation | RMS | Applicable Color Range | QA Plot |
|---|---|---|---|---|
| $i_{ComCam} \to i_{des}$ | $i_{des} - i_{ComCam} = -0.043 (g-i)_{ComCam} +0.017$ | 0.006 | $-0.6 < (g-i)_{ComCam} \leq 1.8$ | link |
| $i_{ComCam} \to i_{des}$ | $i_{des} - i_{ComCam} = -0.122 (g-i)_{ComCam} +0.157$ | 0.009 | $1.8 < (g-i)_{ComCam} \leq 3.7$ | link |
Here are the quality assurance (QA) plots from RTN-099 associated with the first-order piece-wise polynomial fit of the DP1 $i$-band to DES DR2 $i$-band:
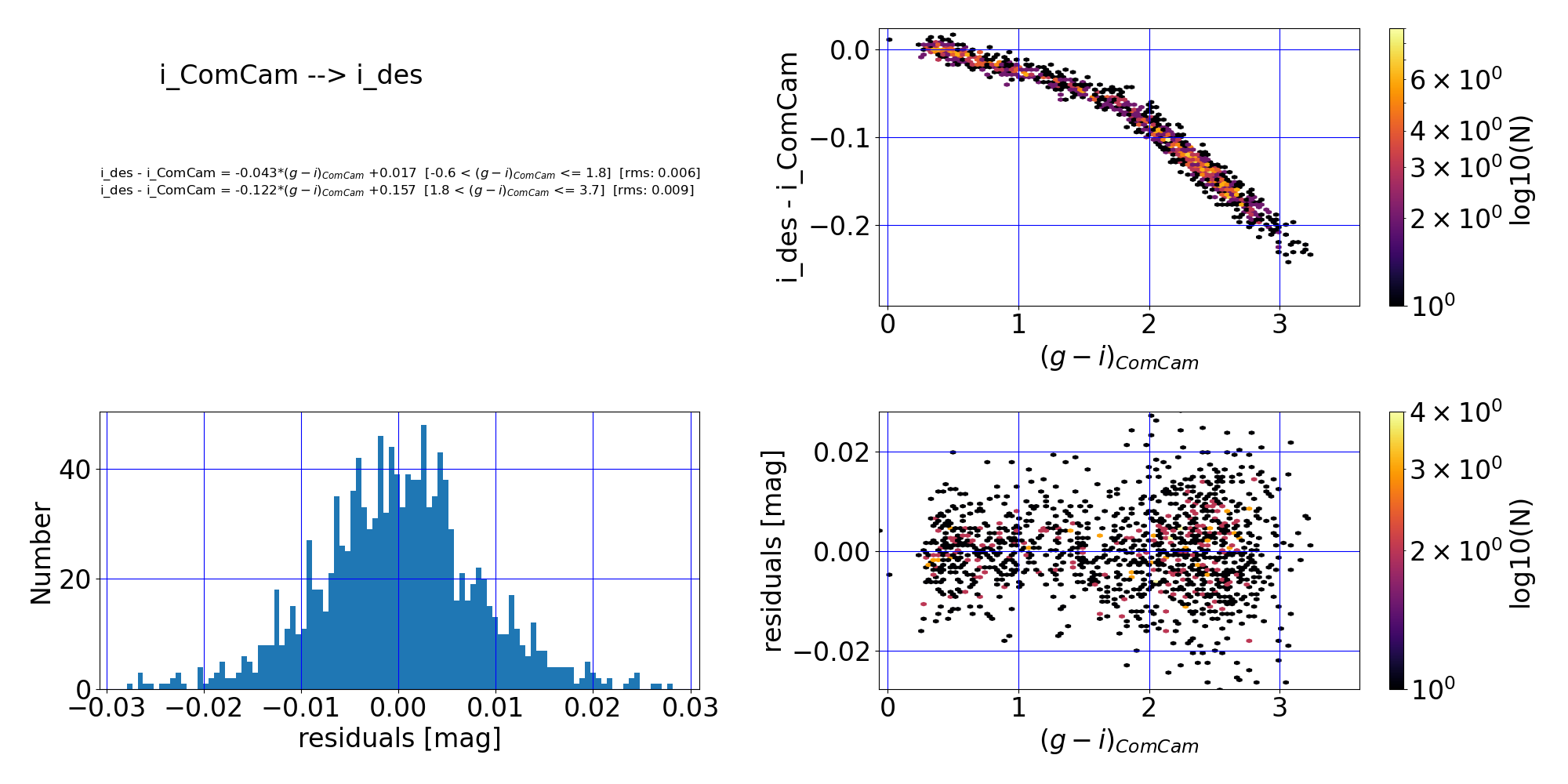
Figure 5: Comparison of LSSTComCam and DES DR2 $i$-band photometry. (Top left:) The fit polynomial equation, the RMS of the fit, and the applicable color range. (Top right:) The difference between the two magnitudes vs. color before any transformation. (Bottom left:) Histogram of the residuals of the fit. (Bottom right:) Residuals of the fit vs. ($g$-$i$) color. The residuals scatter about zero, suggesting that the fit has done a reasonable job.
4.2 Apply the transformation¶
Apply the transformation to the LSSTComCam data in the ECDFS field. This requires first defining the two color ranges in which to apply the separate piece-wise components.
condlist = [
(df_dp1['gi_psfMag'] > -0.6) & (df_dp1['gi_psfMag'] <= 1.8),
(df_dp1['gi_psfMag'] > 1.8) & (df_dp1['gi_psfMag'] <= 3.7)
]
choicelist = [
-0.043 * df_dp1['gi_psfMag'] + 0.017,
-0.122 * df_dp1['gi_psfMag'] + 0.157
]
df_dp1['i_des_poly_offset'] = np.select(condlist, choicelist, default=np.nan)
df_dp1['i_des_poly'] = df_dp1['i_psfMag'] + df_dp1['i_des_poly_offset']
# display(df_dp1)
4.3 Test the transformation¶
Test the transformation by comparing the DES $i$-band magnitudes estimated from the DP1 data in ECDFS against the DES $i$-band magnitudes from the DES DR2 data themselves.
4.3.1 Grab ECDFS data from DES DR2¶
Query the NOIRLab Astro Data Lab TAP service for the DES DR2 data, using suitable constraints for point sources. Note that ADQL cone search does not seem to work for DES DR2; so ranges in ra and dec are used instead.
tap_service = pyvo.dal.TAPService("https://datalab.noirlab.edu/tap")
ra_min = ra_cen - radius/math.cos(math.radians(dec_cen))
ra_max = ra_cen + radius/math.cos(math.radians(dec_cen))
dec_min = dec_cen - radius
dec_max = dec_cen + radius
query = """
SELECT ra, dec,
WAVG_MAG_PSF_G, WAVG_MAG_PSF_R, WAVG_MAG_PSF_I, WAVG_MAG_PSF_Z, WAVG_MAG_PSF_Y,
WAVG_MAGERR_PSF_G, WAVG_MAGERR_PSF_R, WAVG_MAGERR_PSF_I,
WAVG_MAGERR_PSF_Z, WAVG_MAGERR_PSF_Y
FROM des_dr2.main
WHERE (ra BETWEEN %f AND %f) AND (dec BETWEEN %f AND %f) AND
(WAVG_MAGERR_PSF_R BETWEEN 0.00 AND 0.05) AND (WAVG_MAG_PSF_R < 24.0) AND
(EXTENDED_CLASS_WAVG = 0) AND
(FLAGS_G < 4 AND FLAGS_R < 4 AND FLAGS_I < 4)
""" % (ra_min, ra_max, dec_min, dec_max)
results = tap_service.search(query)
table = results.to_table()
df_desdr2 = results.to_table().to_pandas()
# display(df_desdr2)
4.3.2 Match the ECDFS data from LSSTComCam with the ECDFS data from DES DR2¶
Use the cross_match_catalogs function to match the two catalogs.
matches_dp1_desdr2, unmatched_dp1_desdr2 = cross_match_catalogs(df_dp1, df_desdr2,
'coord_ra', 'coord_dec',
'ra', 'dec')
# display(matches_dp1_desdr2)
4.3.3 Compare the LSSTComCam-estimated PS1 i-band magnitudes with the DES DR2 i-band magnitudes¶
Calculate both the differences between the DP1 $i$-band magnitude and the DES DR2 $i$-band magnitude ("before transformation") and the differences between DP1-estimated DES $i$-band magnitude and the DES DR2 $i$-band magnitude ("after transformation"):
matches_dp1_desdr2['dmag_i_cc_des'] = matches_dp1_desdr2['i_psfMag'] -\
matches_dp1_desdr2['match_wavg_mag_psf_i']
matches_dp1_desdr2['dmag_i_des_poly'] = matches_dp1_desdr2['i_des_poly'] -\
matches_dp1_desdr2['match_wavg_mag_psf_i']
Plot the histogram of the $\Delta$mags (both before transformation and post transformation).
dmag_orig = matches_dp1_desdr2.loc[:, 'dmag_i_cc_des']
dmag_new = matches_dp1_desdr2.loc[:, 'dmag_i_des_poly']
dmag_min = -0.3
dmag_max = 0.3
if len(dmag_new) < 100:
plt.hist(dmag_orig, bins=10, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=10, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after transformation')
else:
plt.hist(dmag_orig, bins=100, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=100, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after transformation')
plt.xlabel('$\\Delta$mag [mag]')
plt.ylabel('Number')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"mag difference between LSSTComCam DP1 and DES DR2 $i$-band")
plt.show()
Figure 6: Histogram comparing LSSTComCam and DES DR2 $i$-band photometry for matched stars; differences are calculated as $\Delta$mag = DP1 mag - DES mag. The orange histogram shows the difference before applying the transformation, demonstrating that there is a significant offset in $\Delta$mag values to positive residuals. The blue histogram is the result after applying the polynomial transformation. The transformed difference is centered on a magnitude residual of zero, as expected.
Create a scatter plot of the $\Delta$mags (both before transformation and post transformation) vs. color. Note that the DES DR2 $i$-band differs more from the LSSTCamCom DP1 $i$-band than does the PanSTARRS1 DR2 $i$-band; so the transformation is larger than in the previous example.
color = matches_dp1_desdr2.loc[:, 'gi_psfMag']
dmag_min = -0.3
dmag_max = 0.3
color_min = -0.5
color_max = 3.5
plt.scatter(color, dmag_orig, color=colors[2], alpha=0.50, label='before transformation')
plt.scatter(color, dmag_new, color=colors[5], alpha=0.50, label='after transformation', marker='*')
plt.axis([color_min, color_max, dmag_min, dmag_max])
plt.xlabel('$(g-i)_{ComCam}$')
plt.ylabel('$\\Delta$mag [mag]')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"$\Delta$mag between LSSTComCam DP1 and DES DR2 $i$-band vs. color")
plt.show()
Figure 7: Magnitude differences between LSSTComCam and DES DR2 $i$-band photometry for matched stars plotted as a function of $(g-i)$ color. Orange points show the difference before applying the transformation, demonstrating a large offset in $\Delta$mag that increases at redder $(g-i)$ colors. Blue stars are the results after applying the polynomial transformation. The transformed difference is centered on a magnitude residual of zero, as expected.
5. Second-order polynomial transformation: LSSTComCam $g$ --> Gaia DR3 $BP_{gaia}$¶
Next, look at the polynomial transformation from DP1 $g$-band to Gaia DR3 $BP$-band. Here, a first-order polynomial fit is insufficient, so a second-order polynomial fit is employed. One could consider even higher terms to achieve a more accurate relation, but that is left to the lookup table transformation method described in the next section.
| Conversion | Transformation Equation | RMS | Applicable Color Range | QA Plot |
|---|---|---|---|---|
| $g_{ComCam} \to BP_{gaia}$ | $BP_{gaia} - g_{ComCam} = +0.081 (g-i)_{ComCam}^2 -0.449 (g-i)_{ComCam} +0.162$ | 0.027 | $-0.7 < (g-i)_{ComCam} \leq 3.0$ | link |
Here are the quality assurance (QA) plots from RTN-099 associated with the second-order polynomial fit of the DP1 $g$-band to Gaia DR3 $BP$-band. Note from the bottom-right plot that a third-order term might benefit the fit, but staying with the simpler second-order polynomial still achieves a 0.027 mag RMS in the fit. For more accuracy, using the lookup table method (described in the next section) is recommended.
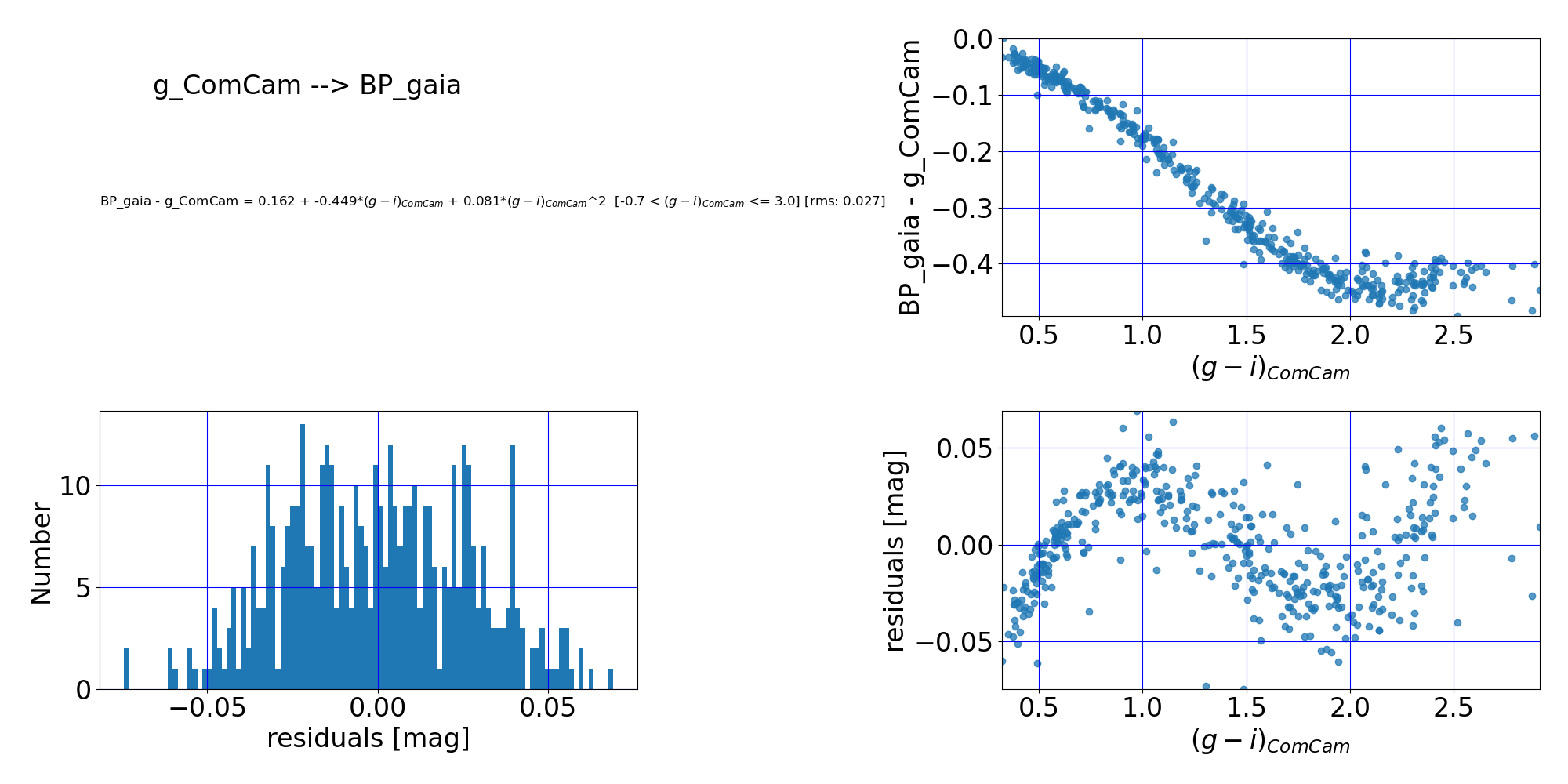
Figure 8: Comparison of LSSTComCam $g$-band and Gaia DR3 $BP$-band photometry. (Top left:) The fit polynomial equation, the RMS of the fit, and the applicable color range. (Top right:) The difference between the two magnitudes vs. color before any transformation. (Bottom left:) Histogram of the residuals of the fit. (Bottom right:) Residuals of the fit vs. ($g$-$i$) color. The residuals scatter about zero, suggesting that the fit has done a reasonable job, though there is a clear pattern remaining that could be mitigated by using a higher-order fit.
5.2 Apply the transformation¶
Select only stars within the applicable color range (-0.7 < $g-i$ < 3.0) from the fit. Apply the polynomial transformation to these stars from the LSSTComCam data in the ECDFS field.
pick_gaia_colors = (df_dp1['gi_psfMag'] < 3.0) & (df_dp1['gi_psfMag'] > -0.7)
df_dp1_gaia_transform = df_dp1[pick_gaia_colors].copy()
df_dp1_gaia_transform['BP_gaia_poly_offset'] = 0.081*np.pow(df_dp1_gaia_transform['gi_psfMag'], 2.) -\
0.449*df_dp1_gaia_transform['gi_psfMag'] + 0.162
df_dp1_gaia_transform['BP_gaia_poly'] = df_dp1_gaia_transform['g_psfMag'] +\
df_dp1_gaia_transform['BP_gaia_poly_offset']
5.3 Test the transformation¶
Test the transformation by comparing the Gaia DR3 $BP$-band magnitudes estimated from the DP1 data in ECDFS against the Gaia DR3 $BP$-band magnitudes from the Gaia DR3 data themselves in ECDFS.
5.3.1 Grab ECDFS data from Gaia DR3¶
Query the Gaia DR3 TAP service, using suitable constraints for a clean sample.
gaia_tap_url = 'https://gaia.aip.de/tap'
gaia_tap = pyvo.dal.TAPService(gaia_tap_url)
query = """
SELECT
source_id, ra, dec, parallax, parallax_error,
pmra, pmra_error, pmdec, pmdec_error,
phot_g_mean_mag, 1.086/phot_g_mean_flux_over_error as phot_g_mean_mag_error,
phot_bp_mean_mag, 1.086/phot_bp_mean_flux_over_error as phot_bp_mean_mag_error,
phot_rp_mean_mag, 1.086/phot_rp_mean_flux_over_error as phot_rp_mean_mag_error,
bp_rp
FROM gaiadr3.gaia_source_lite
WHERE
CONTAINS(POINT('ICRS', ra, dec),CIRCLE('ICRS',%f,%f,%f))=1
AND parallax_over_error > 5
AND ruwe < 1.4
AND phot_g_mean_flux_over_error > 5
AND phot_rp_mean_flux_over_error > 5
AND phot_bp_mean_flux_over_error > 5
""" % (ra_cen, dec_cen, radius)
result = gaia_tap.run_sync(query)
df_gaiadr3 = result.to_table().to_pandas()
# display(df_gaiadr3)
5.3.2 Match the ECDFS data from LSSTComCam with the ECDFS data from Gaia DR3¶
Use the cross_match_catalogs function to match the two catalogs.
matches_dp1_gaiadr3, unmatched_dp1_gaiadr3 = cross_match_catalogs(df_dp1_gaia_transform, df_gaiadr3,
'coord_ra', 'coord_dec',
'ra', 'dec')
# display(matches_dp1_gaiadr3)
5.3.3 Compare the LSSTComCam-estimated BP_gaia magnitudes with the Gaia DR3 BP_gaia magnitudes¶
Calculate both the differences between the DP1 $g$-band magnitude and the Gaia DR3 $BP$-band magnitude ("before transformation") and the differences between DP1-estimated Gaia DR3 $BP$-band magnitude and the actual Gaia DR3 $BP$-band magnitude ("after transformation"):
matches_dp1_gaiadr3['dmag_g_cc_BP_gaia'] = matches_dp1_gaiadr3['g_psfMag'] -\
matches_dp1_gaiadr3['match_phot_bp_mean_mag']
matches_dp1_gaiadr3['dmag_BP_poly'] = matches_dp1_gaiadr3['BP_gaia_poly'] -\
matches_dp1_gaiadr3['match_phot_bp_mean_mag']
Plot the histogram of the $\Delta$mags (both before transformation and post transformation).
dmag_orig = matches_dp1_gaiadr3.loc[:, 'dmag_g_cc_BP_gaia']
dmag_new = matches_dp1_gaiadr3.loc[:, 'dmag_BP_poly']
mag_g_cc = matches_dp1_gaiadr3['g_psfMag']
dmag_min = -0.5
dmag_max = 0.95
if len(dmag_new) < 100:
plt.hist(dmag_orig, bins=10, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=10, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after transformation')
else:
plt.hist(dmag_orig, bins=100, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=100, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after transformation')
plt.xlabel('$\\Delta$mag [mag]')
plt.ylabel('Number')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"difference between LSSTComCam DP1 $g$-band (and transformed $BP$) and Gaia DR3 $BP$-band")
plt.show()
Figure 9: Histogram comparing LSSTComCam $g$-band and Gaia DR3 $BP$-band photometry for matched stars; differences are calculated as $\Delta$mag = DP1 mag - Gaia mag. The orange histogram shows the difference between DP1 $g$ magnitudes and Gaia $BP$ magnitudes before applying the transformation, demonstrating that there is a significant offset in $\Delta$mag values to positive residuals. The blue histogram is the result after applying the polynomial transformation to convert DP1 $g$ magnitudes to Gaia $BP$. The transformed difference is centered on a magnitude residual of zero, as expected.
Create a scatter plot of the $\Delta$mags (both before transformation and post transformation) vs. color. Note that the Gaia DR3 $BP$-band differs substantially from the LSSTCamCom DP1 $g$-band; so the transformation is fairly large.
color = matches_dp1_gaiadr3.loc[:, 'gi_psfMag']
dmag_min = -0.5
dmag_max = 0.95
color_min = -0.5
color_max = 3.5
plt.scatter(color, dmag_orig, color=colors[2], alpha=0.50, label='before transformation')
plt.scatter(color, dmag_new, color=colors[5], alpha=0.50, label='after transformation', marker='*')
plt.axis([color_min, color_max, dmag_min, dmag_max])
plt.xlabel('$(g-i)_{ComCam}$')
plt.ylabel('$\\Delta$mag [mag]')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"$\Delta$mag between LSSTComCam DP1 $g$-band (and transformed $BP$-band) and Gaia DR3 $BP$-band vs. color")
plt.show()
Figure 10: Magnitude differences between LSSTComCam $g$-band and Gaia DR3 $BP$-band photometry for matched stars plotted as a function of $(g-i)$ color. Orange points show the difference between DP1 $g$ magnitudes and Gaia $BP$ magnitudes before applying the transformation, demonstrating a large offset in $\Delta$mag that increases toward redder $(g-i)$ colors. Blue stars are the results after applying the polynomial transformation. The transformed difference is centered on a magnitude residual of zero, as expected.
6. Lookup table transformation: LSSTComCam $g$ --> Gaia DR3 $BP_{gaia}$¶
Next, look at the lookup table transformation from DP1 $g$-band to Gaia DR3 $BP$-band.
Here are the quality‑assurance (QA) plots from RTN-099 that document the lookup‑table transformation from the DP1 $g$‑band to the Gaia DR3 $BP$‑band. The lookup table is constructed by binning matched stars in the ECDFS field by their Rubin DP1 $(g–i)$ color, and then tabulating the median magnitude offset between the Gaia DR3 RP‑band and the Rubin DP1 g‑band for the stars in each color bin.
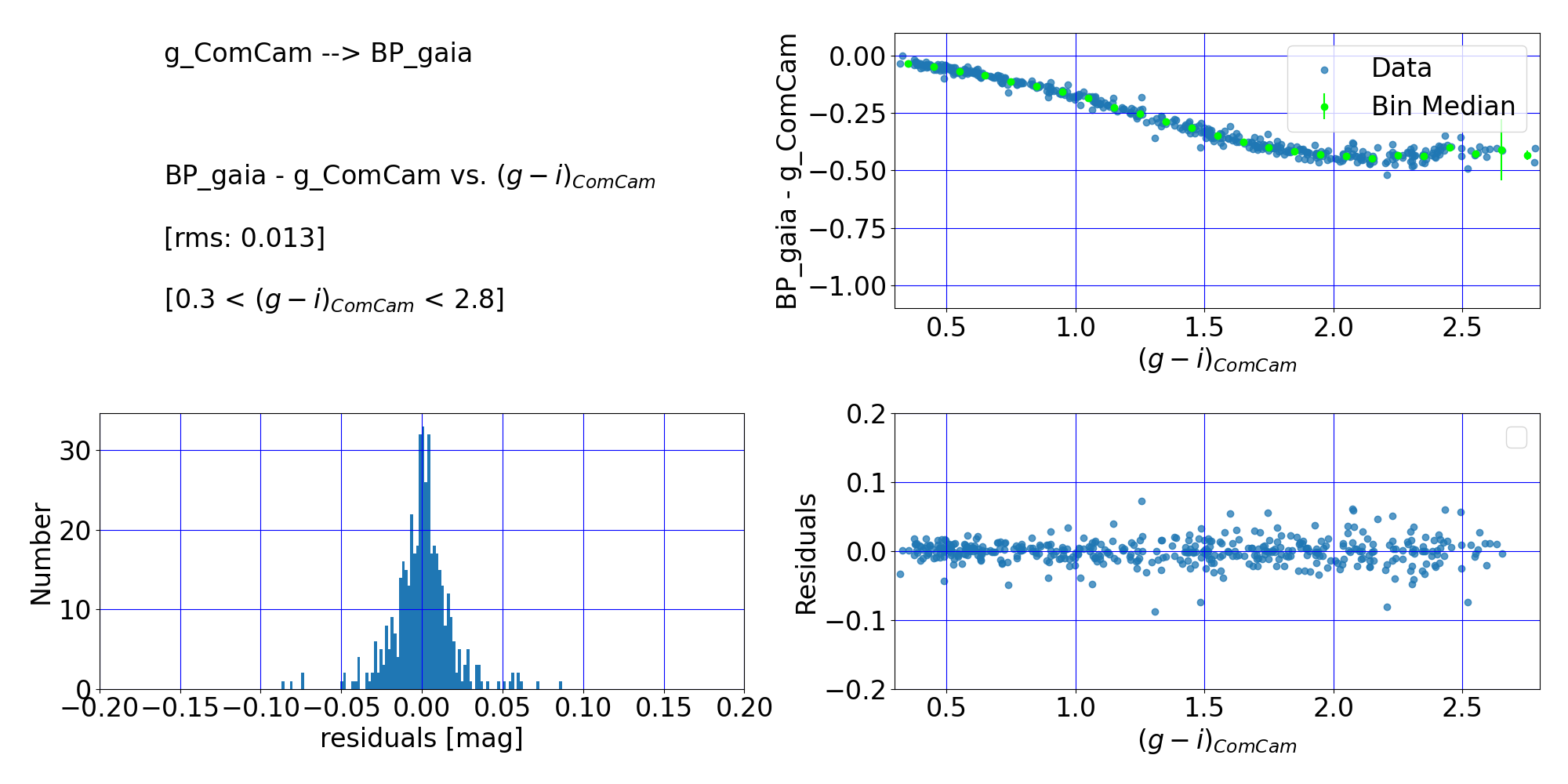
Figure 11: Comparison of LSSTComCam $g$-band and Gaia DR3 $BP$-band photometry. (Top left:) The RMS of the linearly interpolated lookup table relation, and the applicable color range. (Top right:) The difference between the two magnitudes vs. color before any transformation. (Bottom left:) Histogram of the residuals from the interpolated lookup table relation. (Bottom right:) Residuals of the interpolated lookup table relation vs. ($g-i$) color. The residuals scatter about zero, suggesting that the fit has done a reasonable job, and the clear pattern remaining after the polynomial fit in Figure 8 has been removed.
6.2 Apply the transformation¶
Copy the URL of the lookup table link from the RTN-099 line in Section 6.1 and assign it to the variable lut_url:
lut_url = "https://rtn-099.lsst.io/_downloads/54cfc025a795e6ca410bd9dde0f11dda/transInterp.ComCam_to_GaiaDR3_ECDFS.BP_gaia_gi_ComCam.csv"
Read the lookup table over the network into a pandas dataframe, df_lut, and then display its contents.
df_lut = pd.read_csv(lut_url)
df_lut
| bin_label | bin_interval | bin_num | bin_mean | bin_stddev | bin_stderr | bin_median | bin_rstddev | bin_unc | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.35 | (0.3, 0.4] | 13 | -0.031628 | 0.013325 | 0.003847 | -0.032934 | 0.007477 | 0.002599 |
| 1 | 0.45 | (0.4, 0.5] | 32 | -0.049921 | 0.012825 | 0.002303 | -0.049748 | 0.010894 | 0.002414 |
| 2 | 0.55 | (0.5, 0.6] | 36 | -0.065647 | 0.009385 | 0.001586 | -0.066948 | 0.009199 | 0.001922 |
| 3 | 0.65 | (0.6, 0.7] | 23 | -0.082396 | 0.010904 | 0.002325 | -0.085100 | 0.010291 | 0.002689 |
| 4 | 0.75 | (0.7, 0.8] | 23 | -0.104497 | 0.047338 | 0.010092 | -0.113668 | 0.015097 | 0.003945 |
| 5 | 0.85 | (0.8, 0.9] | 15 | -0.136744 | 0.017537 | 0.004687 | -0.132753 | 0.007937 | 0.002569 |
| 6 | 0.95 | (0.9, 1.0] | 20 | -0.158882 | 0.018708 | 0.004292 | -0.157347 | 0.014995 | 0.004202 |
| 7 | 1.05 | (1.0, 1.1] | 24 | -0.189826 | 0.018843 | 0.003929 | -0.183414 | 0.019646 | 0.005026 |
| 8 | 1.15 | (1.1, 1.2] | 13 | -0.220930 | 0.015722 | 0.004538 | -0.224068 | 0.013401 | 0.004658 |
| 9 | 1.25 | (1.2, 1.3] | 16 | -0.250503 | 0.025536 | 0.006593 | -0.251977 | 0.019101 | 0.005985 |
| 10 | 1.35 | (1.3, 1.4] | 12 | -0.291294 | 0.025324 | 0.007636 | -0.287390 | 0.016222 | 0.005869 |
| 11 | 1.45 | (1.4, 1.5] | 24 | -0.318964 | 0.021783 | 0.004542 | -0.314554 | 0.012465 | 0.003189 |
| 12 | 1.55 | (1.5, 1.6] | 25 | -0.347475 | 0.018643 | 0.003806 | -0.347849 | 0.017322 | 0.004342 |
| 13 | 1.65 | (1.6, 1.7] | 16 | -0.369150 | 0.024201 | 0.006249 | -0.375009 | 0.017862 | 0.005597 |
| 14 | 1.75 | (1.7, 1.8] | 27 | -0.395756 | 0.017008 | 0.003335 | -0.399994 | 0.011895 | 0.002869 |
| 15 | 1.85 | (1.8, 1.9] | 18 | -0.417523 | 0.014932 | 0.003622 | -0.416714 | 0.008537 | 0.002522 |
| 16 | 1.95 | (1.9, 2.0] | 19 | -0.431626 | 0.017254 | 0.004067 | -0.430386 | 0.012914 | 0.003713 |
| 17 | 2.05 | (2.0, 2.1] | 18 | -0.430474 | 0.025351 | 0.006149 | -0.437686 | 0.023699 | 0.007001 |
| 18 | 2.15 | (2.1, 2.2] | 16 | -0.446041 | 0.019095 | 0.004930 | -0.449103 | 0.013671 | 0.004284 |
| 19 | 2.25 | (2.2, 2.3] | 17 | -0.442011 | 0.030363 | 0.007591 | -0.433220 | 0.026085 | 0.007929 |
| 20 | 2.35 | (2.3, 2.4] | 22 | -0.439219 | 0.021204 | 0.004627 | -0.436813 | 0.013936 | 0.003724 |
| 21 | 2.45 | (2.4, 2.5] | 14 | -0.400712 | 0.025716 | 0.007132 | -0.400312 | 0.017145 | 0.005743 |
| 22 | 2.55 | (2.5, 2.6] | 8 | -0.430982 | 0.028903 | 0.010924 | -0.427936 | 0.017106 | 0.007580 |
| 23 | 2.65 | (2.6, 2.7] | 4 | -0.688770 | 0.560702 | 0.323721 | -0.411083 | 0.213253 | 0.133637 |
| 24 | 2.75 | (2.7, 2.8] | 2 | -0.434127 | 0.043721 | 0.043721 | -0.434127 | 0.022908 | 0.020302 |
Create a 1-d linear interpolation of the median magnitude offset (bin_median) vs. the midpoint of the DP1 $(g-i)$-color bin (bin_label).
response = interpolate.interp1d(df_lut.bin_label.values.astype(float),
df_lut.bin_median.values,
bounds_error=False, fill_value=0.,
kind='linear')
Calculate and apply the offsets derived from the lookup table. Optionally display the results.
df_dp1_gaia_transform['BP_gaia_lut_offset'] = response(df_dp1_gaia_transform['gi_psfMag'].values)
df_dp1_gaia_transform['BP_gaia_lut'] = df_dp1_gaia_transform['g_psfMag'] +\
df_dp1_gaia_transform['BP_gaia_lut_offset']
# display(df_dp1)
6.3 Test the transformation¶
Test the transformation by comparing the Gaia DR3 $BP$-band magnitudes estimated from the DP1 data in ECDFS against the Gaia DR3 $BP$-band magnitudes from the Gaia DR3 data themselves.
6.3.1 Grab ECDFS data from Gaia DR3¶
This was already done in Section 5.3.1. No need to do it again.
6.3.2 Match the ECDFS data from LSSTComCam with the ECDFS data from Gaia DR3¶
We will re-do this, as df_dp1 has additional information now.
matches_dp1_gaiadr3, unmatched_dp1_gaiadr3 = cross_match_catalogs(df_dp1_gaia_transform, df_gaiadr3,
'coord_ra', 'coord_dec',
'ra', 'dec')
# display(matches_dp1_gaiadr3)
6.3.3 Compare the LSSTComCam-estimated BP_gaia magnitudes with the Gaia DR3 BP_gaia magnitudes¶
Calculate differences between DP1-estimated Gaia DR3 $BP$-band magnitude from the lookup table method and the actual Gaia DR3 $BP$-band magnitude ("after transformation"): We re-do this, since there is new info from the lookup table method.
matches_dp1_gaiadr3['dmag_BP_lut'] = matches_dp1_gaiadr3['BP_gaia_lut'] -\
matches_dp1_gaiadr3['match_phot_bp_mean_mag']
Plot the histogram of the $\Delta$mags before transformation, after the polynomial-based transformation, and after the lookup-table-based transformation.
dmag_new_lut = matches_dp1_gaiadr3.loc[:, 'dmag_BP_lut']
dmag_min = -0.5
dmag_max = 0.95
if len(dmag_new) < 100:
plt.hist(dmag_orig, bins=10, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=10, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after polynomial transformation')
plt.hist(dmag_new_lut, bins=10, range=(dmag_min, dmag_max), color=colors[3],
alpha=0.50, label='after LUT transformation')
else:
plt.hist(dmag_orig, bins=100, range=(dmag_min, dmag_max), color=colors[2],
alpha=0.50, label='before transformation')
plt.hist(dmag_new, bins=100, range=(dmag_min, dmag_max), color=colors[5],
alpha=0.50, label='after polynomial transformation')
plt.hist(dmag_new_lut, bins=100, range=(dmag_min, dmag_max), color=colors[3],
alpha=0.50, label='after LUT transformation')
plt.xlabel('$\\Delta$mag [mag]')
plt.ylabel('Number')
plt.grid(True)
plt.grid(color='grey')
plt.legend(loc='upper left', fontsize=8)
plt.title(r"mag difference between LSSTComCam DP1 $g$-band (and transformed $BP$-band) and Gaia DR3 $BP$-band")
plt.show()
Figure 12: Histogram comparing LSSTComCam $g$-band and Gaia DR3 $BP$-band photometry for matched stars; differences are calculated as $\Delta$mag = DP1 mag - Gaia mag. The orange histogram shows the difference between DP1 $g$ magnitudes and Gaia $BP$ magnitudes before applying the transformation, demonstrating that there is a significant offset in $\Delta$mag values to positive residuals. The blue histogram is the result after applying the polynomial transformation to convert DP1 $g$ magnitudes to Gaia $BP$. The pink histogram shows the result after applying the lookup table correction, providing an even narrower distribution about zero net residual.
Create a scatter plot of the $\Delta$mags vs. color for before transformation, for after the polynomial-based transformation, and for after the lookup-table-based transformation. Note the small systematic trends vs. color seen in the the polynomial-based transformation are effectively eliminated by the lookup-table-based transformation.
color = matches_dp1_gaiadr3.loc[:, 'gi_psfMag']
dmag_min = -0.35
dmag_max = 0.85
color_min = -0.5
color_max = 3.5
plt.scatter(color, dmag_orig, color=colors[2], alpha=0.50,
label='before transformation')
plt.scatter(color, dmag_new, color=colors[5], alpha=0.50,
label='after polynomial transformation', marker='*')
plt.scatter(color, dmag_new_lut, color=colors[3], alpha=0.75,
label='after LUT transformation', marker='s', facecolors='none')
plt.axis([color_min, color_max, dmag_min, dmag_max])
plt.xlabel('$(g-i)_{ComCam}$')
plt.ylabel('$\\Delta$mag [mag]')
plt.grid(True)
plt.grid(color='grey')
plt.legend()
plt.title(r"$\Delta$mag between LSSTComCam DP1 $g$-band (and transformed $BP$-band) and Gaia DR3 $BP$-band vs. color")
Text(0.5, 1.0, '$\\Delta$mag between LSSTComCam DP1 $g$-band (and transformed $BP$-band) and Gaia DR3 $BP$-band vs. color')
Figure 13: Magnitude differences between LSSTComCam $g$-band and Gaia DR3 $BP$-band photometry for matched stars plotted as a function of $(g-i)$ color. Orange points show the difference between DP1 $g$ magnitudes and Gaia $BP$ magnitudes before applying the transformation, demonstrating a large offset in $\Delta$mag that increases toward redder $(g-i)$ colors. Blue stars are the results after applying the polynomial transformation. Pink open squares show the residuals after transforming using the lookup table, demonstrating smaller residuals than the polynomial results.